library(tidyverse)
df <- read_csv("https://raw.githubusercontent.com/yesols/aou-workshop/refs/heads/main/data/final_dataset.csv")
head(df)Exploratory Data Analysis and Visualization
1 Learning Objectives
In this session, we will learn common functions to explore data and visualize data using the synthetic dataset we worked with last time.
At the end of Session 3, you will be able to:
- Use R commands to understand data.
- Perform descriptive statistics.
- Visualize data with box plots, histograms, and scatter plots.
2 Setup
We will begin by loading packages and reading in a file.
# A tibble: 6 × 10
person_id year_of_birth race sex_at_birth condition_concept_id
<dbl> <dbl> <chr> <chr> <dbl>
1 1 1987 Unknown Male NA
2 2 1987 Asian Female NA
3 3 2004 Asian Male NA
4 4 1944 Black Male NA
5 5 1941 Asian Male NA
6 6 2006 Other Male NA
# ℹ 5 more variables: condition_start_date <date>,
# condition_source_value <chr>, drug_concept_id <dbl>,
# drug_exposure_start_date <date>, SBP <dbl>This file is what we saved after merging dataframes in the previous session, but I added an additional column SBP for systolic blood pressure.
3 Create Necessary Variables for Analysis
We will create an age column from year_of_birth.
df <- df %>% mutate(
age = 2026 - year_of_birth
)
head(df)# A tibble: 6 × 11
person_id year_of_birth race sex_at_birth condition_concept_id
<dbl> <dbl> <chr> <chr> <dbl>
1 1 1987 Unknown Male NA
2 2 1987 Asian Female NA
3 3 2004 Asian Male NA
4 4 1944 Black Male NA
5 5 1941 Asian Male NA
6 6 2006 Other Male NA
# ℹ 6 more variables: condition_start_date <date>,
# condition_source_value <chr>, drug_concept_id <dbl>,
# drug_exposure_start_date <date>, SBP <dbl>, age <dbl>It is also useful to have binary variables to indicate their disease status and drug exposure status. We will use if_else() function for this purpose. if_else() takes three arguments: condition to evaluate, value if the evaluation is true, value if false.
df <- df %>%
mutate(
condition_status = if_else(is.na(condition_concept_id), 0, 1), # 0 if condition_concept_id is NA, 1 if not
exposure_status = if_else(is.na(drug_concept_id), 0, 1) # 0 if drug_concept_id is NA, 1 if not
)4 Contingency Table
Let’s take a quick look at MCI/dementia cases breakdown by sex.
df %>% count(sex_at_birth, condition_status)# A tibble: 6 × 3
sex_at_birth condition_status n
<chr> <dbl> <int>
1 Female 0 28
2 Female 1 16
3 Male 0 37
4 Male 1 11
5 Other 0 5
6 Other 1 3janitor package has a nice cross-tabulation function:
library(janitor)
df %>% tabyl(sex_at_birth, condition_status) sex_at_birth 0 1
Female 28 16
Male 37 11
Other 5 35 Summarize Data
summary() function can give you a quick overview of data and its distribution. It also will show you a count of NAs in each column. You can use summary() on a vector (or a single column from a dataframe) or the entire dataframe.
summary(df$SBP) Min. 1st Qu. Median Mean 3rd Qu. Max. NA's
84.0 108.0 119.0 117.2 127.0 149.0 5 summary(df) person_id year_of_birth race sex_at_birth
Min. : 1.00 Min. :1936 Length:100 Length:100
1st Qu.: 25.75 1st Qu.:1957 Class :character Class :character
Median : 50.50 Median :1972 Mode :character Mode :character
Mean : 50.50 Mean :1973
3rd Qu.: 75.25 3rd Qu.:1990
Max. :100.00 Max. :2007
condition_concept_id condition_start_date condition_source_value
Min. :4128031 Min. :1988-02-15 Length:100
1st Qu.:4128031 1st Qu.:2016-04-30 Class :character
Median :4128031 Median :2026-12-22 Mode :character
Mean :4151509 Mean :2028-10-20
3rd Qu.:4182210 3rd Qu.:2041-07-23
Max. :4182210 Max. :2071-04-19
NA's :70 NA's :70
drug_concept_id drug_exposure_start_date SBP age
Min. :19073183 Min. :1987-09-16 Min. : 84.0 Min. :19.00
1st Qu.:19073183 1st Qu.:2013-02-12 1st Qu.:108.0 1st Qu.:35.75
Median :19073183 Median :2025-05-28 Median :119.0 Median :54.50
Mean :19073183 Mean :2029-02-16 Mean :117.2 Mean :52.91
3rd Qu.:19073183 3rd Qu.:2043-04-05 3rd Qu.:127.0 3rd Qu.:69.00
Max. :19073183 Max. :2080-10-17 Max. :149.0 Max. :90.00
NA's :65 NA's :65 NA's :5
condition_status exposure_status
Min. :0.0 Min. :0.00
1st Qu.:0.0 1st Qu.:0.00
Median :0.0 Median :0.00
Mean :0.3 Mean :0.35
3rd Qu.:1.0 3rd Qu.:1.00
Max. :1.0 Max. :1.00
Grouped summaries using group_by() and summarise() can be helpful.
df %>%
group_by(sex_at_birth) %>%
summarise(
n = n(),
mean_SBP = mean(SBP, na.rm = TRUE),
sd_SBP = sd(SBP, na.rm = TRUE),
median_age = median(age, na.rm = TRUE)
)# A tibble: 3 × 5
sex_at_birth n mean_SBP sd_SBP median_age
<chr> <int> <dbl> <dbl> <dbl>
1 Female 44 116. 16.6 51.5
2 Male 48 118. 13.1 56.5
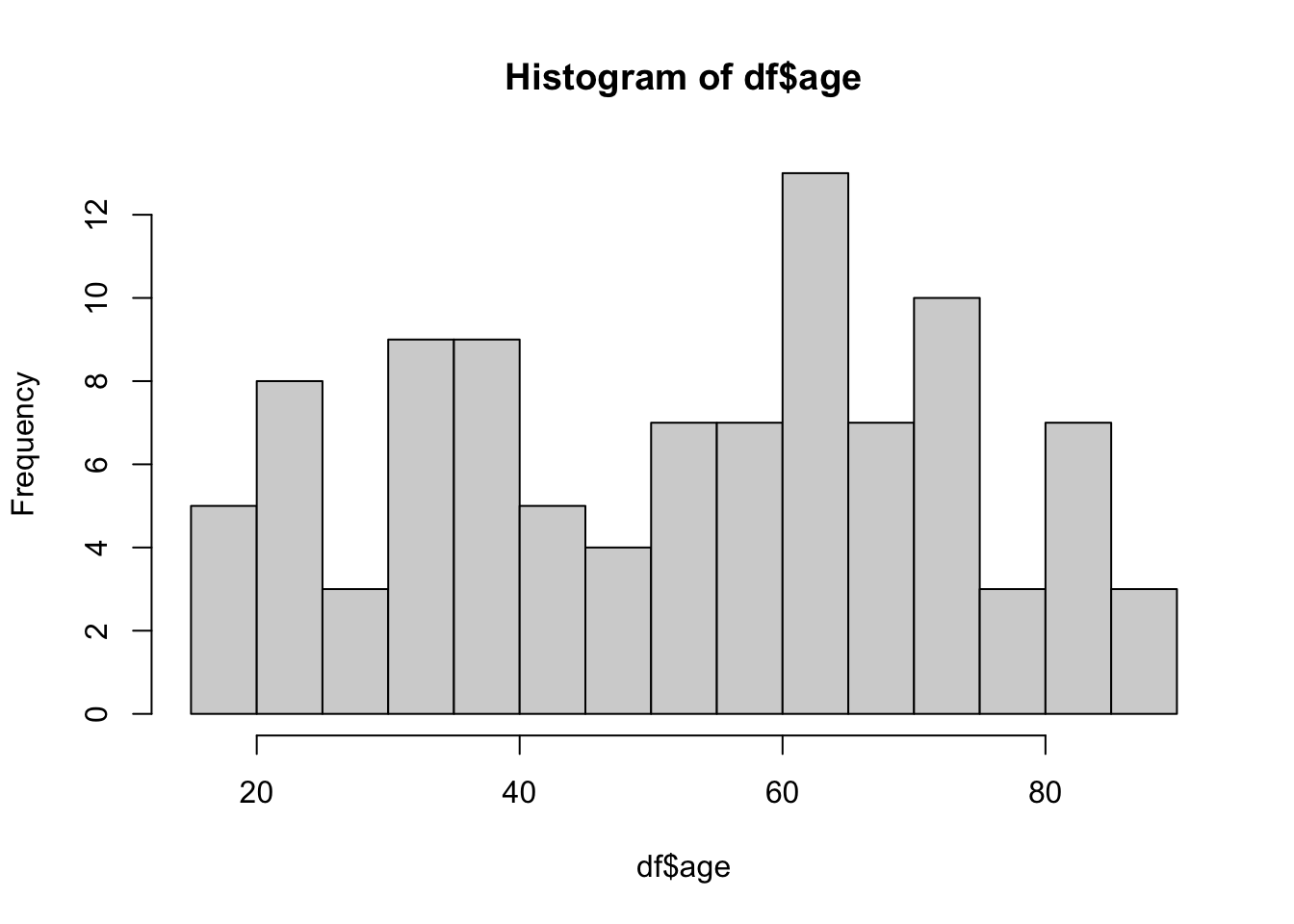
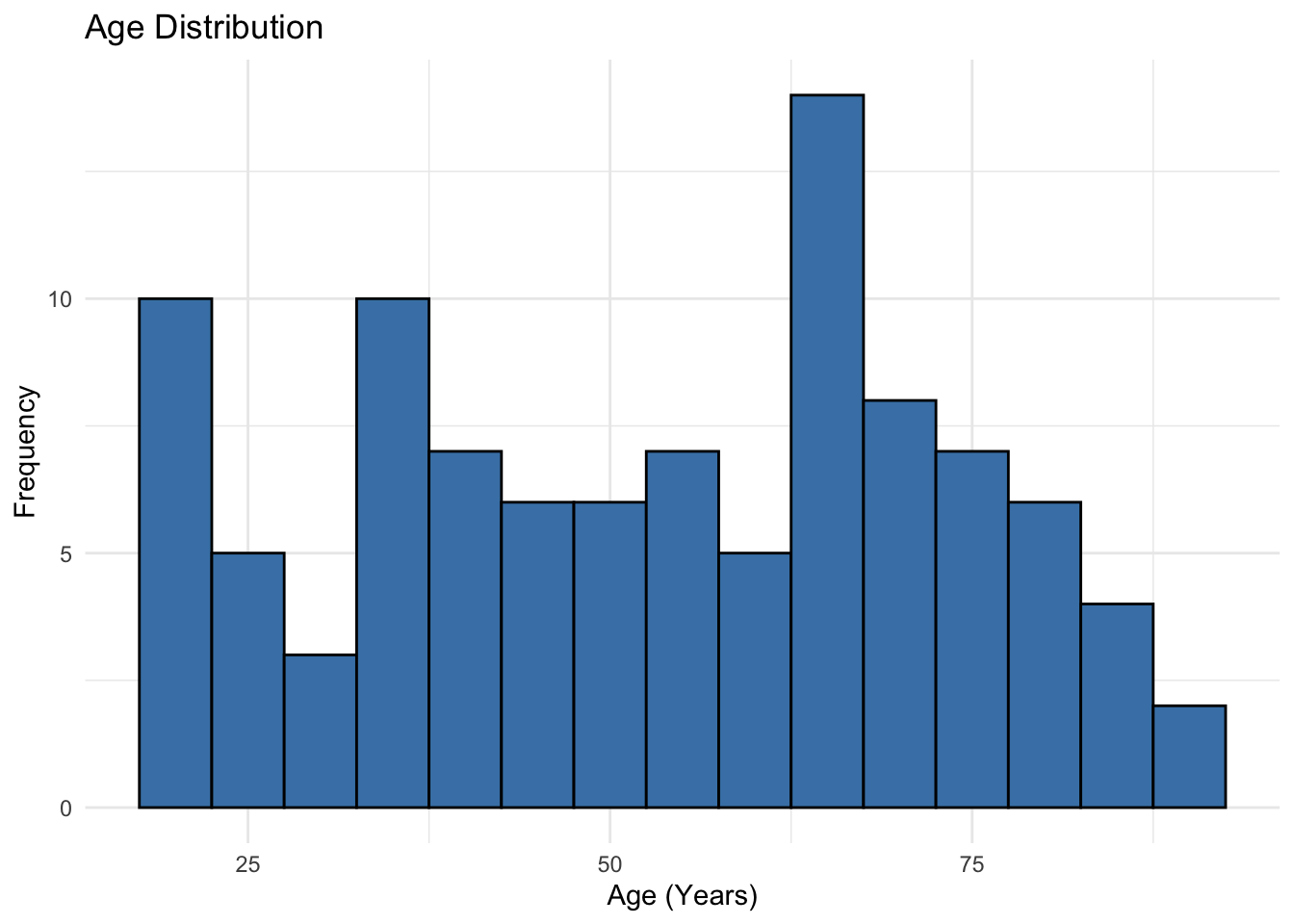
3 Other 8 117. 9.05 59.56 Histogram
Histogram is a good way to visualize distribution of continuous data. I like base R commands for a quick visualization on the fly. But ggplot() from ggplot2 package (included already when we loaded tidyverse) is highly customizable and recommended for publication-quality plots. Below, you can see the code for both.
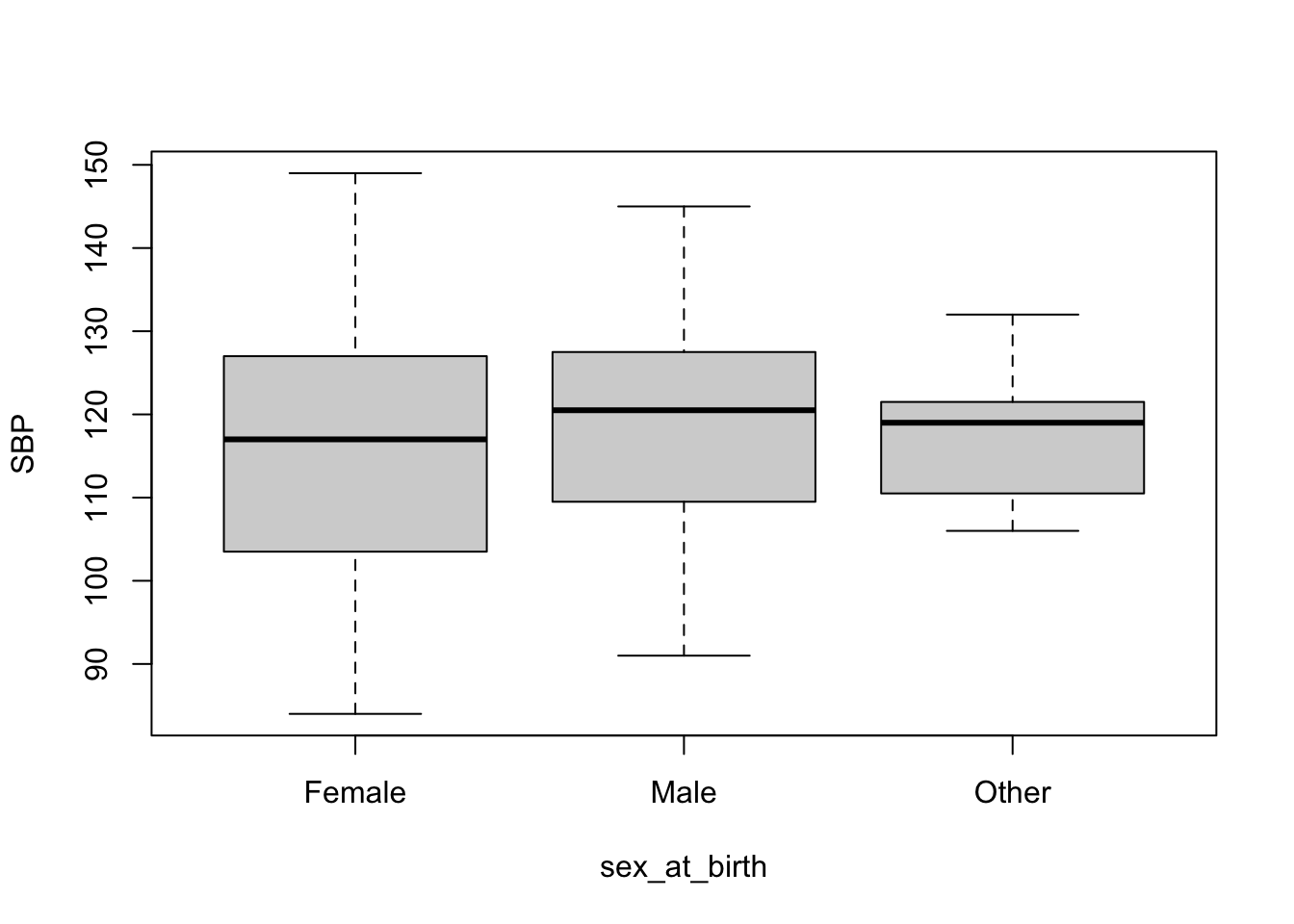
7 Box Plot
Box plots are good for visualizing continuous data broken down to some groups. Here, we will visualize distribution of SBP for each sex_at_birth category. Again, I will show both base R version and ggplot2 version.
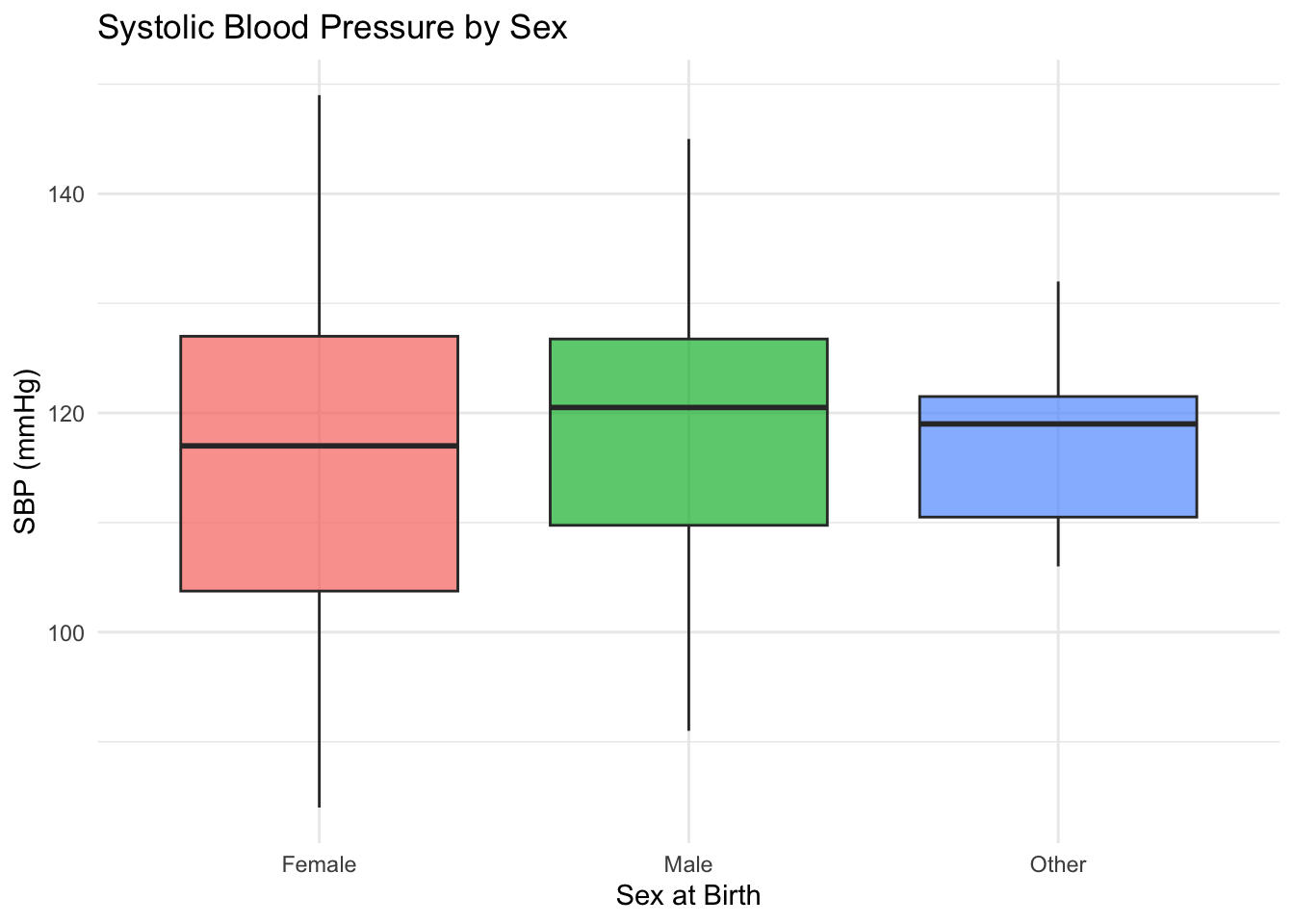
# ggplot2 box plot
ggplot(df, aes(x = sex_at_birth, y = SBP, fill = sex_at_birth)) +
geom_boxplot(alpha = 0.7) +
labs(
title = "Systolic Blood Pressure by Sex",
x = "Sex at Birth",
y = "SBP (mmHg)"
) +
theme_minimal() +
theme(legend.position = "none") # Suppress redundant legendWarning: Removed 5 rows containing non-finite outside the scale range
(`stat_boxplot()`).Take a note of the warning message in ggplot2 version that says 5 rows were removed. This often occurs when there are NAs in the data. We know SBP column had NA values. If this was unexpected, you need to inspect the data and handle the out-of-range values accordingly.
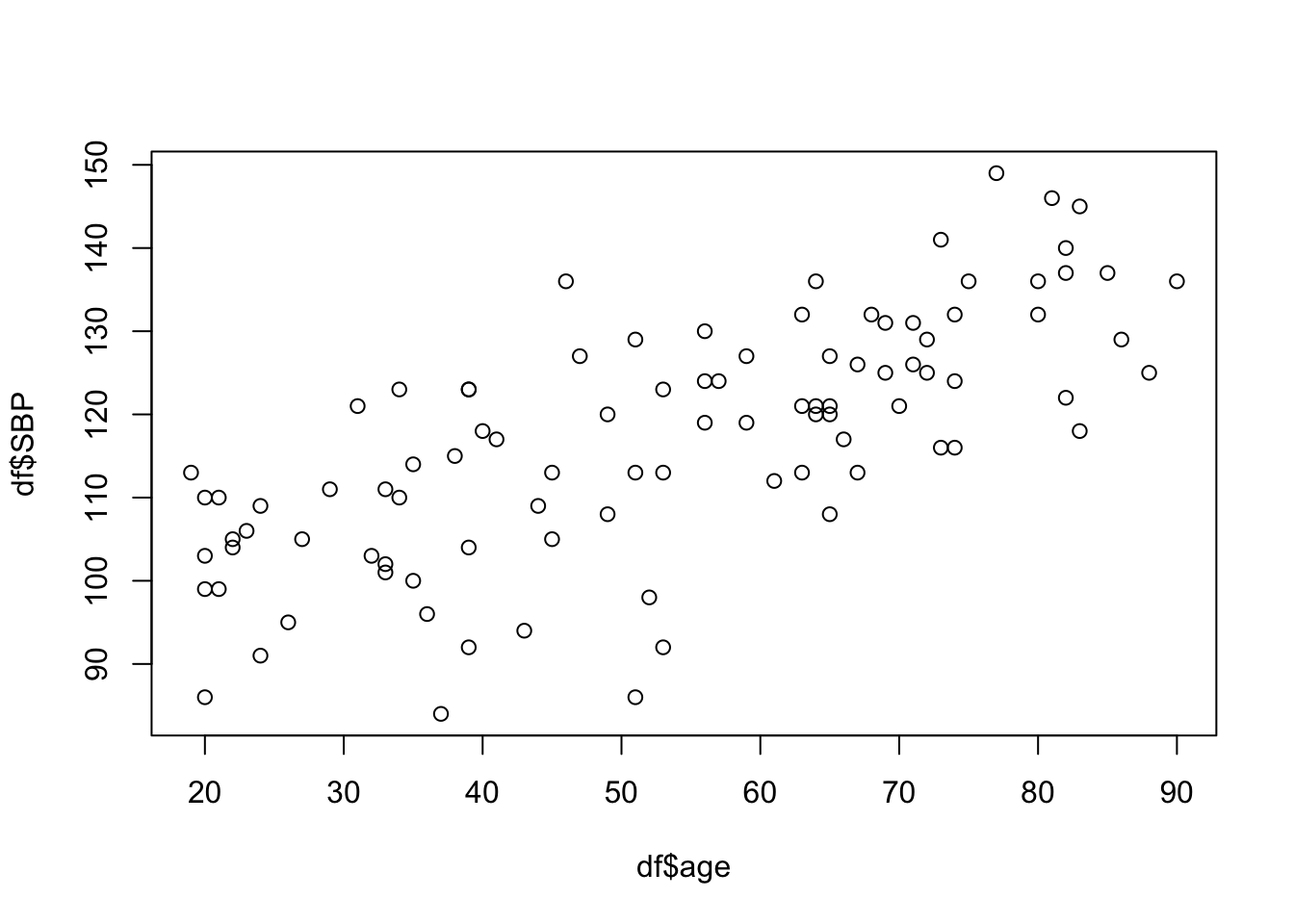
8 Scatter Plot
We use scatter plots for visualizing the relationship between two continuous variables. We will plot SBP against age.
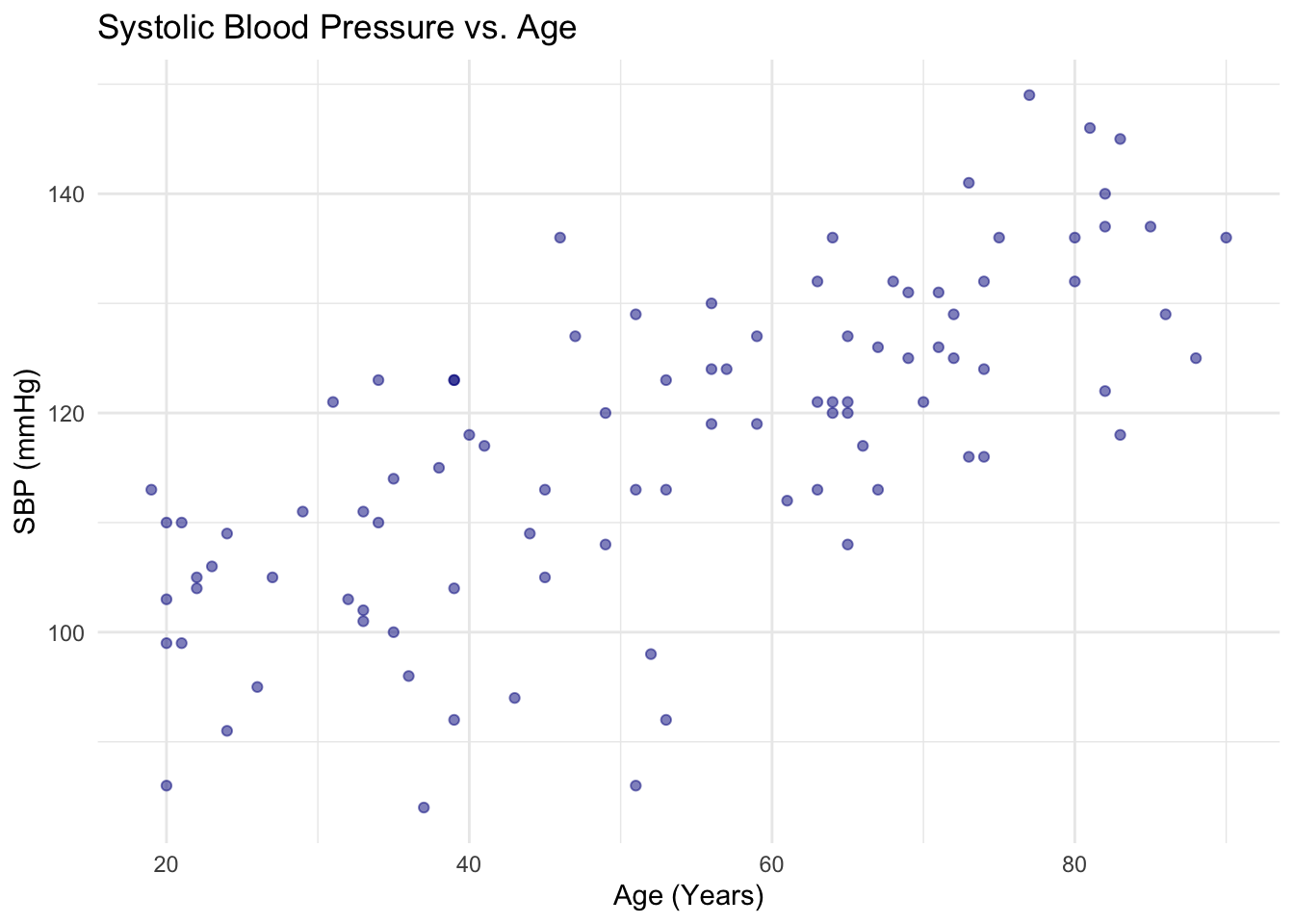
# ggplot2 scatter plot
ggplot(df, aes(x = age, y = SBP)) +
geom_point(color = "darkblue", alpha = 0.5) +
labs(
title = "Systolic Blood Pressure vs. Age",
x = "Age (Years)",
y = "SBP (mmHg)"
) +
theme_minimal()Warning: Removed 5 rows containing missing values or values outside the scale range
(`geom_point()`).9 Preparing for Next Session
In session 4, we will learn how to perform basic statistical tests in R. Because this workshop is focused on learning the technical aspect of programming, discussions of statistical concepts will be minimal. Review the following methods as needed prior to the next session:
- t-test and ANOVA
- chi-square test
- linear regression
- logistic regression (WWAMI Research Methods course does not cover this. So we will go over this concept in a little more detail than others.)
Also, we will apply the graphical and statistical functions on the real All of Us data. So, if you have not already, ensure you have a dataset prepared for your own project by next session. It does not have to be perfect - constructing a dataset always takes a long time. Just have a dataset that has demographic variables and an outcome variable at a minimum.